IHES
IHES
Robert Penner’s recent findings could help to predict effective antiviral targets, more resistant to mutation
Robert Penner, René Thom Chair at IHES, has continued his study of SARS CoV-2, the virus that causes covid. In recent work [1], he analyzed the English, South African and Brazilian mutations of concern that are currently spreading worldwide, among others. His methods allow the prediction of so-called backbone free energy (BFE), which is a measure of the springlike energy potential associated with a certain part of a protein. He has shown that BFE can help predict regions primed for rearrangement, based upon the geometry of a protein alone. The method was applied to the SARS CoV-2 spike glycoprotein, which is the target of all the vaccines thus far approved for France or the US, with particular attention to the protein regions that have mutated in the variants of concern, which are now predominant in many parts of France for example.
There are several key findings, including that 11 of the 19 mutated amino acid residues (18 of them illustrated in the figure) are either free from hydrogen bonds or lie in disorganized regions of the protein, giving a clue on where to expect future mutations. This is critical to the further development of vaccines, as we want them to remain effective as viruses continue to mutate, in particular against SARS CoV-2, and in general against the “next virus X”. Another novelty of this work is the prediction of likely function for several of the 8 other mutated amino acids as inferred from the behavior of BFE with varying acidity, a technique only recently feasible due to availability of protein structures uncovered using experimental breakthroughs in Cryo Electron Microscopy, which can now image proteins such as the spike at near-atomic resolution at different acidities.
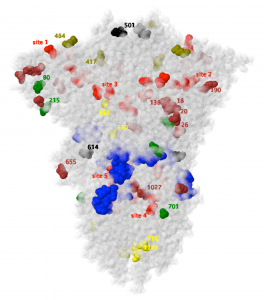

Penner’s earlier work had identified 5 so-called sites of interest in the spike glycoprotein as vaccine or drug targets that were promising due to large BFE conserved across all human coronavirus spikes (illustrated in red in the figure). However, these sites cannot be targeted using the mRNA or virus-vector methods of the current vaccines based on nucleic acids because the geometry of high BFE regions is too labile to survive display and response by the immune system. Corresponding nearby passive sites of interest of low BFE are reported (illustrated in blue in the figure) in the current work that do provide promising targets for nucleic acid vaccines. Moreover, Penner argues that these sites are unlikely to mutate going forward due to lack of evolutionary pressures, a takeaway that again applies not only to SARS CoV-2 but also to virus X. Another reason that targeted sites such as these are crucial is that increased disease morbidity indicates the insufficiency of native immune response to the full spike. Science must be sneakier than Nature in this case, whether for future covid or for the disease caused by virus X.
Penner is in communication with colleagues at Institut Pasteur and with the vaccine development team at Moderna who have both helped steer the current work.
[1] Penner, Robert. “Antiviral Resistance against Viral Mutation: Praxis and Policy for SARS-CoV-2” Computational and Mathematical Biophysics, vol. 9, no. 1, 2021, pp. 81-89. https://doi.org/10.1515/cmb-2020-0119